Experimental Evaluation of PROTACs
Experimental Aims 1 and 2 endeavour to produce a suite of PROTAC ligands that target ATM and POP1 for degradation. Aim 3 will test the action of these therapeutics, and determine optimal dosage, through initial in vitro studies, then further analysis in an in vivo mouse model.
Cell Line Preparation
In vitro studies of the PROTAC ligands will be conducted in a mammary carcinoma cell line called MFM-233 from Sigma Aldrich. These are cells that were obtained from a post-menopausal patient’s grade III metastatic breast cancer tissue (1). Cells will be handled in accordance with the ECACC guidelines, whereby they are cultured in MEM + 2 mM Glutamine + 10% Foetal Bovine Serum (FBS). This culture requires physical containment level 2 (2).
It is anticipated that approximately 36 putative PROTAC ligands will be produced by my colleagues. Each will be analysed as part of the in vitro studies, including the telomerase assay, western blot, and flow cytometry. Increasing concentrations of each PROTAC ligand (0, 1, 2, 3, 4, 5, 10, 15, 20 µg/mL) will be incubated alongside MFM-233 cells for 24 hours (3). A positive control will be analysed, involving cell treatment with a well-established telomerase inhibitor, MST-312 (4). The best candidates from these results will be selected for further study in a mouse model.
Terminal Restriction Fragment Analysis
Terminal restriction fragment analysis will be used to quantify the length of telomeres of MFM-233 cell cultures treated with the various PROTAC designs. We will use a modified method as reported by (5). Untreated MFM-233 cells will be used as the negative control, with MFM-233 cells treated with the telomerase inhibitor MST-312 as the positive control (4). We anticipate that the application of our designed PROTACs will significantly decrease the telomere length via degradation of essential proteins within the telomere complex.
Methods
The DNA of the treated and untreated cells must first be extracted. This will be done with a PureLink Genomic DNA Mini Kit per the manufacturer’s instructions. This will then be analysed for the DNA concentration with a spectrophotometer, aiming for about 100 ug/mL. If under this amount, the DNA should be precipitated with ethanol to further concentrate the solution.
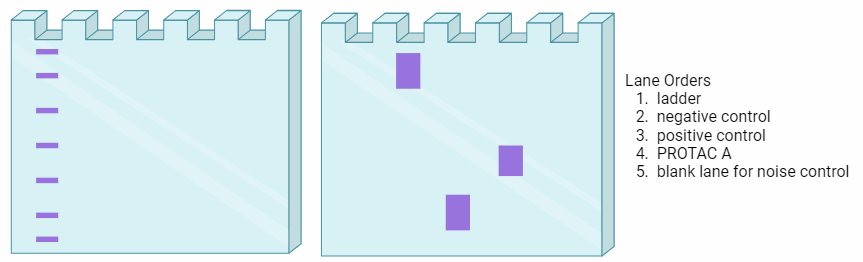
Next, a DNA integrity assessment should be performed. This is done via gel electrophoresis with ethidium bromide for nucleic acid imaging. When imaged with UV light, DNA with good integrity should have distinct, high molecular weight bands. These samples are suitable to proceed with the Southern Blot.
These suitable samples should then be digested with the specific restriction enzymes, RsaI and HinfI, and incubated for a minimum of 16 hrs. The digested DNA should then be processed with gel electrophoresis. It is important to leave empty lanes to use for background noise comparison.
The first step in imaging is to lay a filter paper flat on top of the finished gels, and letting the gel dry. This will then be transferred to sit in an amount of water that covers the gel, the filter paper will be removed and the gel will incubate for a short period. These steps increase the structural stability of the gel so that it can be easily manipulated.
The gel will then be rolled within a nylon mesh, and placed into a hybridization tube. The gel will then be stained using SSC solution and green fluorescent nucleic acid stain in a dark place. The gel will then be removed, washed and imaged with a phosphorimager at 520 BP, 40 Blue and 488 nm. Following this, the gel will be placed in denaturing solution for a short period of time and then washed. This is repeated with neutralisation solution, and then the gel can be again placed in a hybridization tube with hybridization solution and incubated for a similar amount of time as before.
Finally, the radioactive telomere probe will be prepared by combining 2 ng of telomere probe with polynucleotide kinase reaction buffer, polynucleotide kinase and γ32P-dATP. After centrifugation, a quick incubation and centrifugation again, the radioactive probe will be loaded into a G-25 chromatography microspin column and centrifuged again. The filtrate, which holds the probe, will then be removed and placed in the hybridization tube with the gel for a second incubation period.
Once removed, the gel will be incubated once in SCC solution and incubated twice with SCC and SDS. This is now ready for imaging again with the phosphorimager (5).
Predicted Results
Western Blot
Western blotting will be used to quantify the relative abundance of our target proteins in response to PROTAC-induced degradation (6). Untreated cells (negative control) will be compared with cells incubated in increasing concentrations of the PROTAC ligands (0, 1, 2, 3, 4, 5, 10, 15, 20 µg/mL) designed by the BCMB team. We anticipate that this aspect of the experiment will quantify whether the PROTAC ligand has effectively induced POP1 and ATM degradation in the cell, as well as reveal optimal concentrations at which the therapy will be effective.
Methods
Cells will be lysed, then undergo a series of centrifugation and buffer suspension steps (7). Protein concentrations will be quantified using a BCA assay kit from Thermofisher Scientific. Samples will be diluted to give uniform protein concentration with 2 x Laemmli sample buffer prior to being loaded into a 12% SDS-PAGE gel. Samples will be loaded alongside Precision Plus Protein WesternC Standards from BioRad. Following electrophoresis, proteins will be transferred onto a polyvinyldene digluoride (PVDF) membrane (6).
The membrane will be washed with 1 x TBST, then non-specific binding will be blocked with blocking buffer (5% (w/v) skim milk powder in TBST) at room temperature overnight. The next day, following a series of washing steps, membranes will be incubated with the primary antibody of interest (rabbit polyclonal anti-POP1, 1:16000 and mouse monoclonal anti-GAPDH, 1:100000, or rabbit polyclonal anti-ATM, 1:2000 and mouse monoclonal anti-GAPDH, 1:100000). More washing steps will proceed. Secondary antibody solution (HRP-conjugated Affinipure Goat Anti-Mouse IgG(H+L), 1:10000 and HRP-conjugated Affinipure Goat Anti-Rabbit IgG(H+L), 1:10000) will be added and left for incubation. The membrane will again be washed thrice. Target proteins will be visualised with SuperSignal West Pico PLUS Chemiluminescent Substrate from ThermoFisher Scientific. Membranes will be visualised with a ChemiDoc machine (6).
Predicted Results
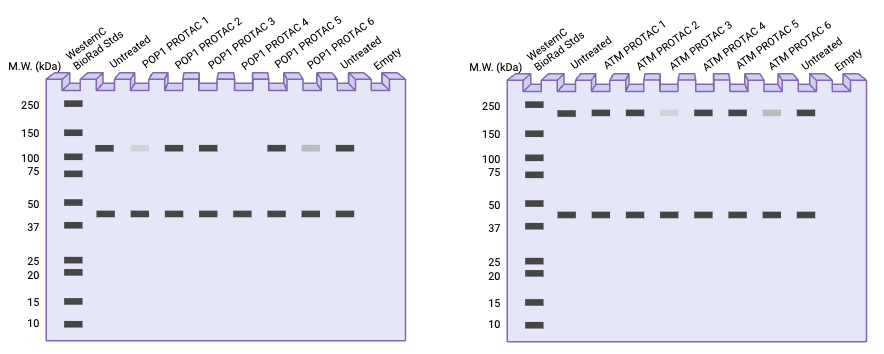
Analysis
Figure 2 shows a predicted westen blot result for both POP1 and ATM. It is hypothesised that some PROTAC ligands will be more successful than others in terms of target protein knockdown. The most successful ligands will result in a reduction in abundance of the protein, seen as a loss of density in the band at the protein’s molecular weight. The ligands and concentrations that show the largest reduction in protein abundance signal will be selected for further study in the animal model.
Flow Cytometry
Flow cytometry will be used to detect the proportion of cells in each PROTAC-treated sample undergoing cell death in comparison of untreated samples. The preferential induction of cell death (apoptosis, necrosis, etc.) is a key goal of cancer therapeutics (8–11). By knocking down proteins involved in the recruitment (ATM) and stabilisation (POP1) of the telomerase complex, we hope to specifically induce cell death in tumour cells, while leaving the host’s non-cancerous cells intact. We anticipate that the results of this assay will allow our team to assess the successful induction of early and late apoptosis in our cell cultures in response to PROTAC treatment. A successful PROTAC system will show a higher proportion of cells in the early and late stages of apoptosis compared to untreated control samples.
An Annexin V flow cytometry assay will be performed to quantify the progression of cell death in MFM-233 cells treated with increasing concentrations of our PROTAC therapeutic (0, 1, 2, 3, 4, 5, 10, 15, 20 µg/mL) (12). In healthy viable cells, aminophosholipids like phosphatidylserine (PS) are present only on the inner layer of the lipid bilayer membrane. During the early stages of apoptosis, the plasma membrane is disrupted, and PS becomes exposed on the outer side of the membrane (7). Annexin V conjugated with APC binds PS on the outer side of membranes, and is therefore a good indicator of early apoptosis occurring in a cell that can be detected by flow cytometry (12). Propium iodide (PI) is a fluorescent dye that binds DNA. When a cell is in the late stages of apoptosis, it’s membrane becomes permeable to this dye, and it will bind intracellular DNA. PI cannot pass through a healthy cell membrane (13)
Methods
Cell suspensions will be collected into Falcon tubes, then centrifuged at 150 x g for 5 minutes. Cells will be washed in PBS (12). At least three technical replicates will need to be performed for each sample. Cells will be resuspended in 10X binding buffer (AVBB) from ThermoFisher Scientific. Fluorochrome-conjugated Annexin V will be added to the solution, then it will incubate at room temperature for 10 minutes. Cells will then be re-suspended in 1X binding buffer. 7-AAD Viability Staining Solution will then be added and left to incubate at room temperature for 10 minutes. Samples will then be analysed by flow cytometry as per the manufacturer’s instructions (7).
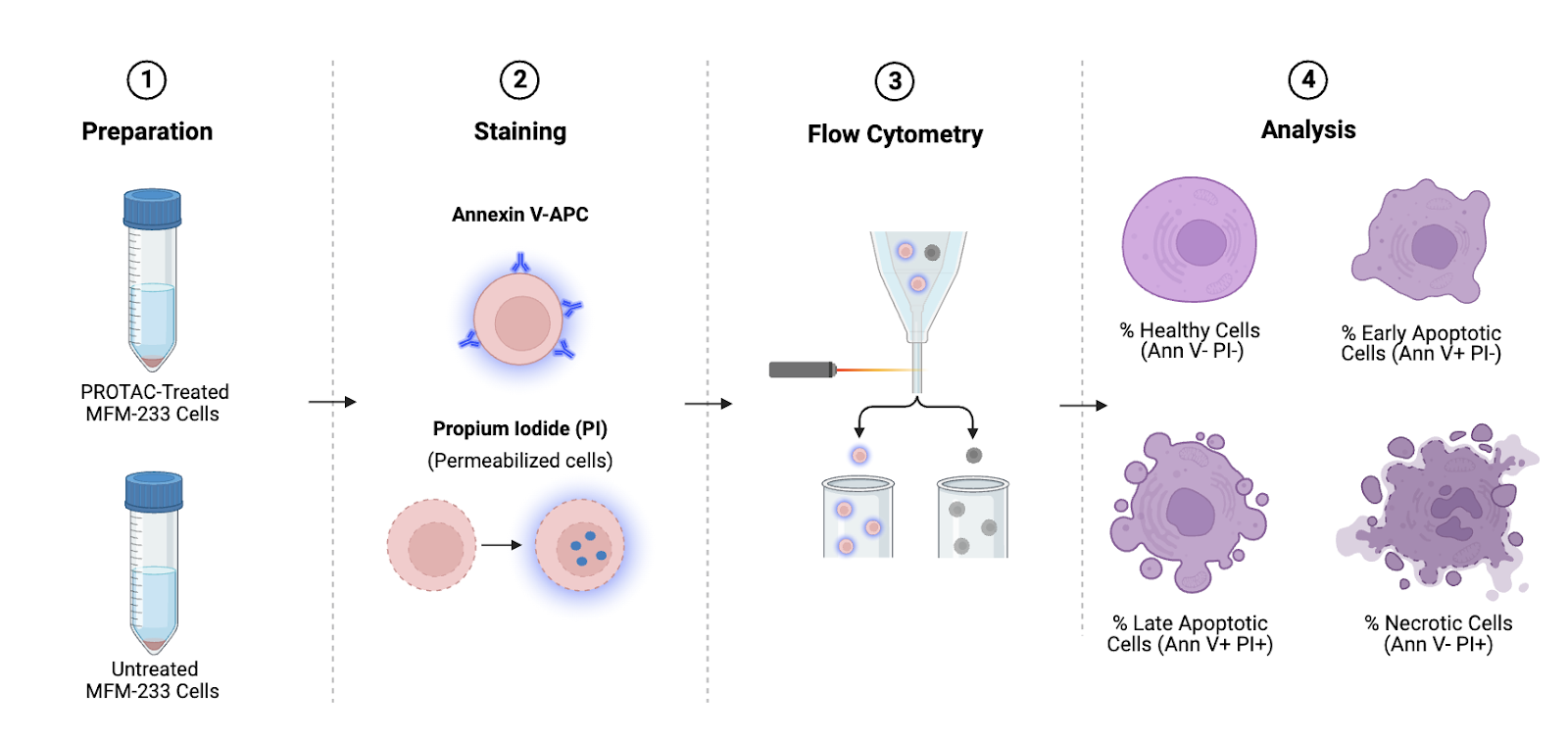
Analysis
This assay will indicate the successful induction of cell death in MFM-233 cells in response to our PROTAC therapeutics. Viable, healthy cells will negatively atain for both Annexin V and PI. Early apoptotic cells will stain as Annexin-V positive but PI negative. Cells in late apoptosis will stain positive for both Annexin V and PI. Necrotic cells will stain positive for PI but negative for Annexin V (12). At least three biological replicates will be conducted for each treatment group, and these results will be averaged across them.
Expected Outcomes
This aspect of the experiment will demonstrate which PROTAC ligand and concentration best induces cell death in MFM-233 cells. It is expected that successful therapeutics will show a greater proportion of cells undergoing oncolysis, compared to untreated control cells. The results of these assays will inform the PROTAC ligands and relevant dosages that will be used for the following mouse model studies.
Mouse Model Preparation
Initial in vivo studies will be conducted in murine models. By studying tumour progression, volume, and any changes in life span in response to our PROTAC therapies, it is anticipated that this will allow insight into their therapeutic applications in humans.
NOD SCID gamma (NSG) mice are inbred and severely immunocompromised, are optimal for the study of tumours and metastasis in animal models (14,15). These animals are highly susceptible to widespread metastasis and have been effectively used in similar studies (14,16,17). We hypothesise that animals treated with our PROTAC therapeutic will exhibit lower spread of primary tumours, show lower tumour volume, and have extended life-spans as a result.
Methods
MDA-MB-231 cell culture will be injected into our mice to promote breast cancer tumour formation. This cell line was established from a patient’s metastatic mammary adenocarcinoma (18). It has an aggressive, triple-negative and poorly-differentiated phenotype, and is commonly used in the study of breast cancer metastasis (2,19). Prior to introduction to mice, cells will be cultured at 37 ˚C in Leibovitz’s L-15 medium with 2 mM glutamine and 15% foetal bovine serum (FBS), as per the supplier’s instructions (2).
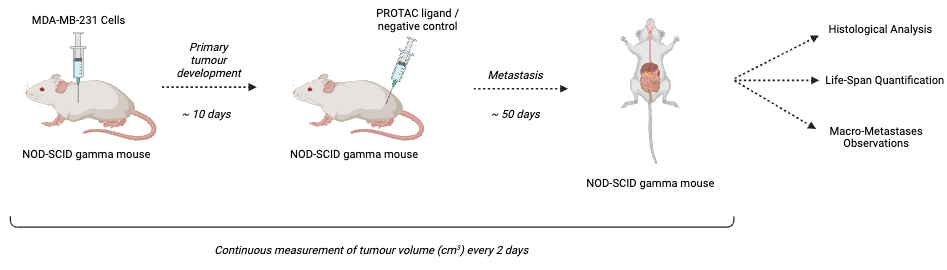
MDA-MB-231 cells will be injected into the mammary fat bads of our NSG mice. Ten days following, the mice will be assessed for the development of primary tumours (14). Tumour volumes will be calculated using the following formula:
\[ \text{Tumour Volume} = \frac{\text{length} \, \cdot \, (\text{width})^2}{2} \]
Next, mice bearing tumours will be randomly allocated to treatment groups. Age, size, and primary tumour volume will remain consistent and controlled. Mice in each group will be either left as a negative control, or treated with increasing concentrations of our putative PROTAC ligand therapeutics. At least ten biological replicates will be used for each treatment group (20). The concentration of PROTAC administered will be informed by the results of the previous in vitro studies. Tumour volume will be consistently assessed over approximately 50 days (14,21). The life-span of these mice will also be taken into account for the analysis of each PROTAC’s efficacy.
Analysis
In addition to tumour volume and life-span, we can observe the development of metastasis post-mortem. Histological analysis of tumours and qualitative observations of macro-metastasis of the tumour into each organ are also likely to give insight into the therapeutic action of our PROTACs (14).
Expected Outcomes
It is anticipated that in vivo studies of our PROTAC ligands will provide a better representation of their therapeutic action in humans, compared to the preliminary studies conducted in cell lines (11). Successful inhibitors of ATM and POP1 are expected to inhibit the spread of primary breast cancer tumours. Successful PROTAC candidates will increase mouse life-span, decrease tumour volume, and quaitatively show metastasis to fewer organs.