Mathematical Validation
The list of potential protein candidates are compiled from the ranked list of proteins from the betweenness method of proteins 2 steps away from telomerase and the top 30 ranked proteins from each resolution parameter of the diffusion distance method.
Essential Proteins
Comparing the list of ‘Essential Proteins’ provided on Ed with our list of potential proteins then removing any essential proteins.
- 1/3 of the proteins from the betweenness method were essential
- 32% of the proteins from the diffusion distance method were essential
Date and Party Hubs (Pearson’s Correlation Coefficient)
Hubs are nodes with high degree, in the context of PPINs, they are proteins which interact with many other proteins. Biologic networks, including PPINs, are particularly susceptible to hub removal. (1)
However, not all hubs are created equal. Removal of ‘date hubs’, which interact with their neighbours at different times or locations, is more likely to be lethal to the cell. (2) Alternatively, removal of ‘party hubs’, which interact with their neighbours at the same time and location is less likely to be lethal to the cell. (2)
Han et. al. propose that date hubs connect multiple functional modules together which party hubs are important nodes within functional modules. If we consider the communities we found in our WalkTrap network partition to be representative of functional modules we can then use the date/party distinction to choose nodes which have the desired deleterious effect on telomerase activity.
To reduce chance of cell lethality we prefer party hubs to date hubs and preferentially party hubs within the telomerase community as this is the target community. However, within the telomerase community we aim to cause fairly serious disruption and so we would prefer proteins which are date hubs within telomerase. Taking this together we arrive at the following decision tree for the potential protein list:
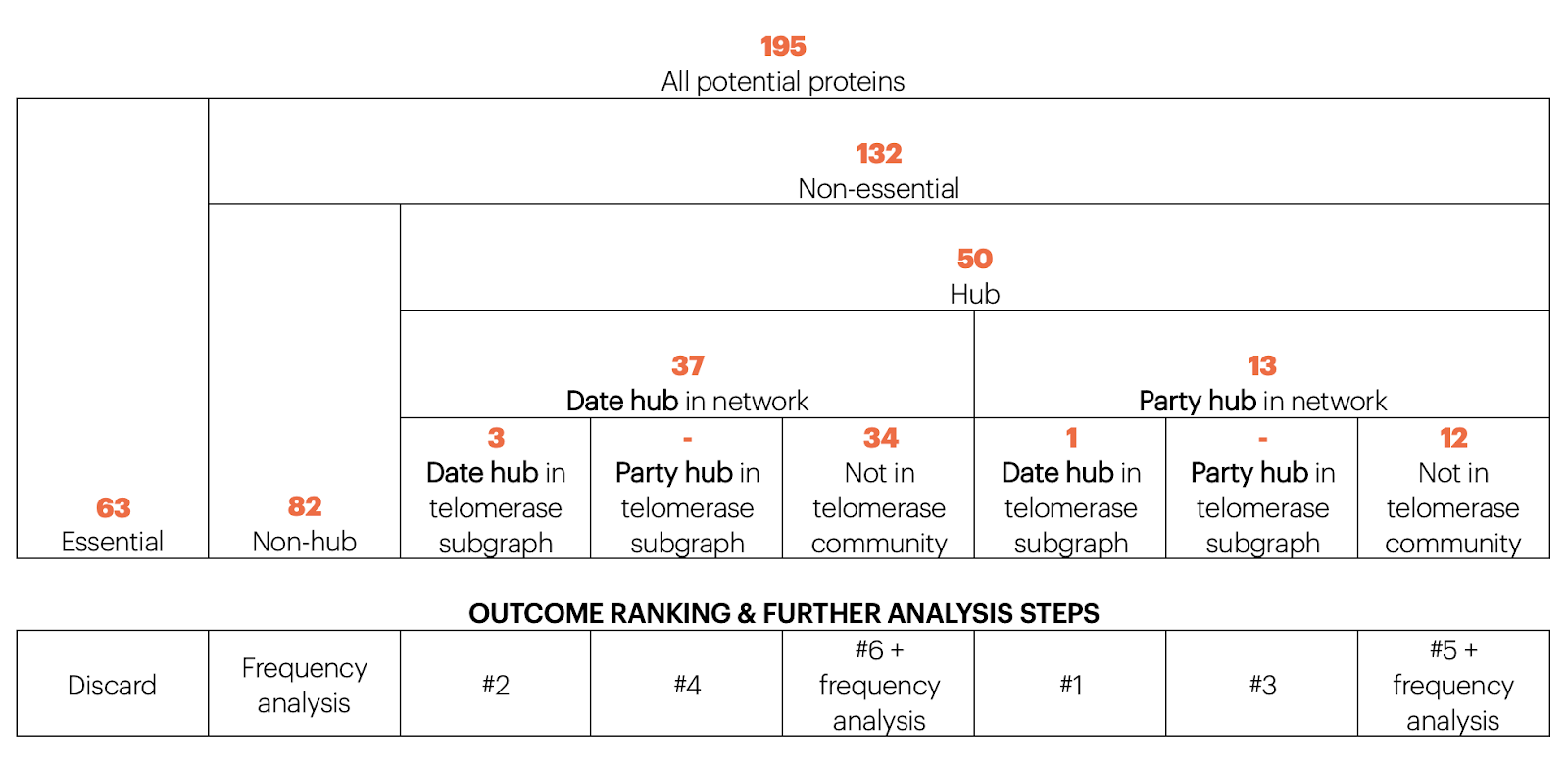
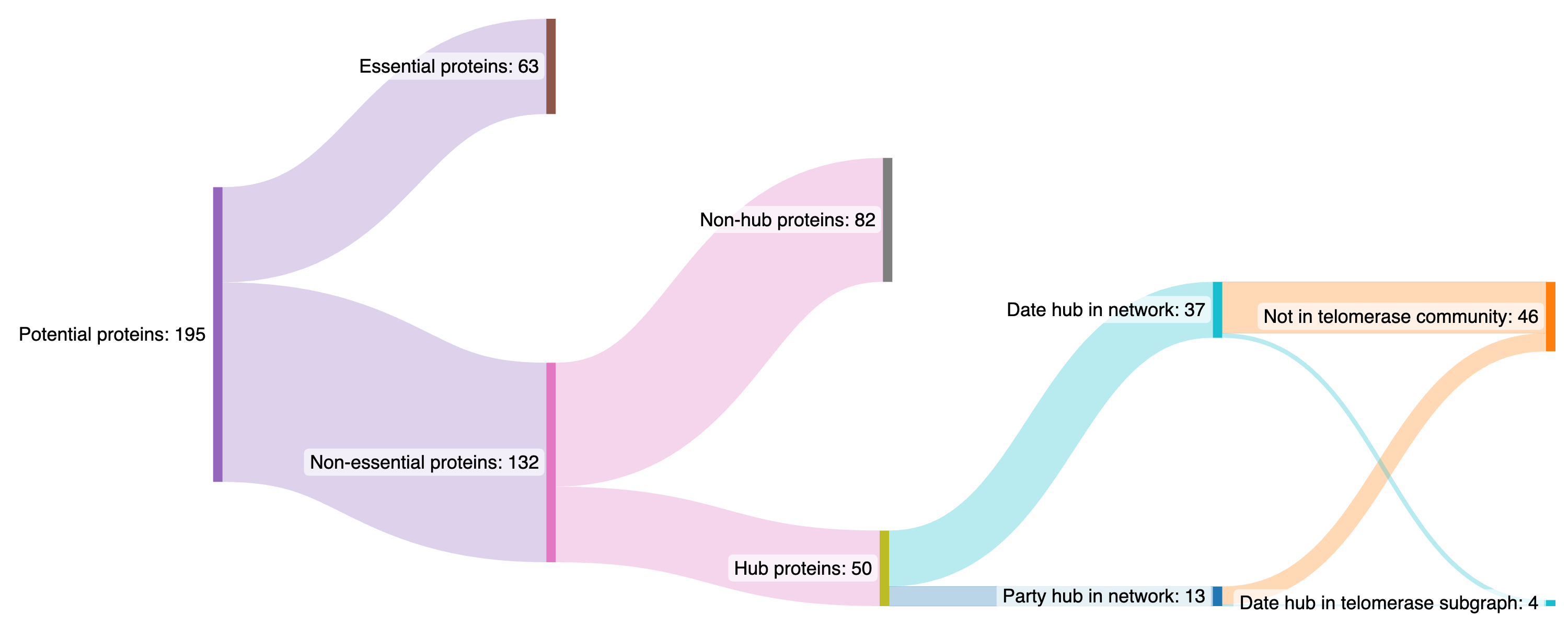
All non-essential proteins produced by the betweenness method were hubs so for the non-hub analysis as there are multiple repetitions we choose those proteins which re-appear most frequently in multiple resolution parameters.
Differentiating Date Hubs from Party Hubs with Pearson’s Correlation Coefficient
Similarity
Similarity is a measure of equivalence between two nodes. One framework for this idea is structural equivalence, which is a measure of how many shared neighbours between each node normalised by various means. Han et. al. uses the similarity measure of Pearson’s correlation coefficient.
Pearson’s Correlation Coefficient
PCC uses the following measure of covariance: \text{cov}(A_i, A_j) = \sum_k(A_{ik}-\langle A_i \rangle)(A_{jk}-\langle A_j \rangle)
Where \langle A_i \rangle is the average of the i th row.
PCC is given as:
r_{ij} = \frac{\text{cov}(A_i, A_j)}{\text{cov}(A_i, A_i)\text{cov}(A_j, A_j)}
Average PCC
Average PCC is a value given to each hub node in the network. It is the average value of r_{ij} between the node i and each of its neighbours j.
Average PCC of hubs in PPINs show a bimodal distribution, date hubs are those hubs contained in the lower cluster, and party hubs those in the higher cluster of average PCC. That is, date hubs show low similarity to their neighbours, this means their neighbours are not highly connected to each other. On the other hand, party hubs have high similarity to their neighbours, meaning their neighbours have a higher degree of interconnection.
The biological implications of average PCC become clear here, date hubs are connecting disparate modules that are less connected to each other whereas party hubs are a part of a more highly connected section of network, more likely to be a functional module.
Implementation
The first step is to determine what defines a hub. Han et. al., who uses a highly cleaned version of the Saccharomyces cerevisiae PPIN, defines a hub as any node with degree greater than 5. The mean node of our network is ~36. Using a series of different definitions of hubs, calculating the avgPCC of all hubs in the network a PCC cutoff of 0.005 was determined.
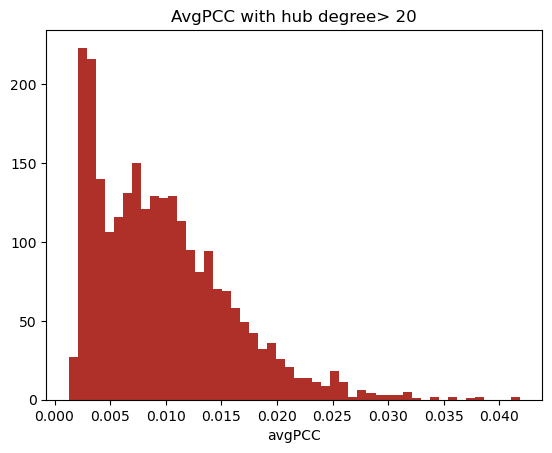
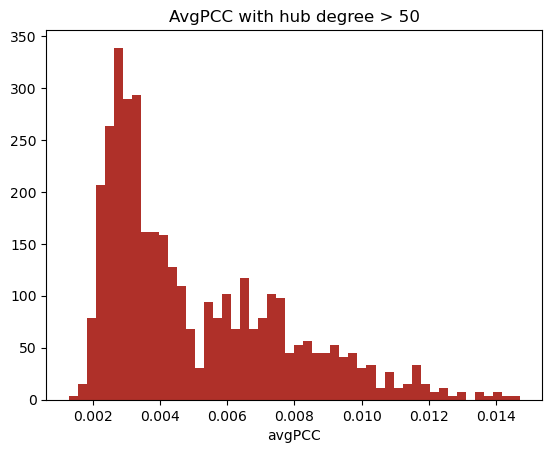
Non-hubs do not display a bimodal distribution of avgPCC (2) which was seen in our network with a hub definition as low as 10 degrees, however we define a hub as a node with degree greater than 30, as these nodes can clearly be defined as hubs. This was also the measure used to segregate non-hubs during initial stages of validation.
A subgraph of the telomerase community was created and avgPCC calculated, this time giving a PCC cutoff of 0.013.
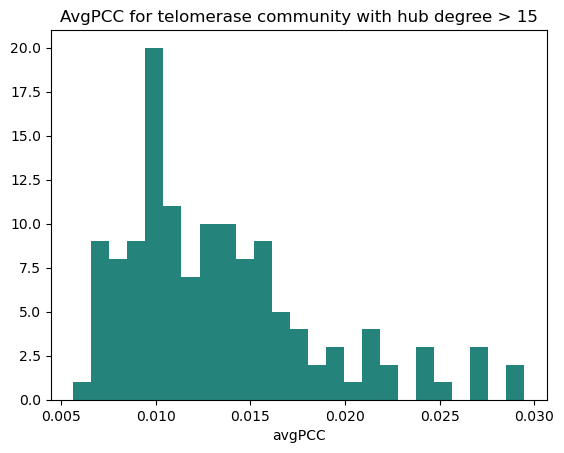
AvgPCC of each protein of interest was then calculated both for the whole network as well as for within the telomerase community if the protein was in it, and they were categorised accordingly.
Non-hub protein frequency analysis
The highest ranked 10 proteins of each resolution parameter from the diffusion distance results were inspected and the more recurring proteins were put forward as potential targets for the biologists to validate.
Results
- Non-hubs: PRP5, ROK1, RPL34A
- Network party hub & telomerase date hub: MRE11
- Network date hub & telomerase date hub: RAD51, RAD52, SGS1
- Diffusion distance & date hub: RPL34A
- Diffusion distance & party hub: EHD3